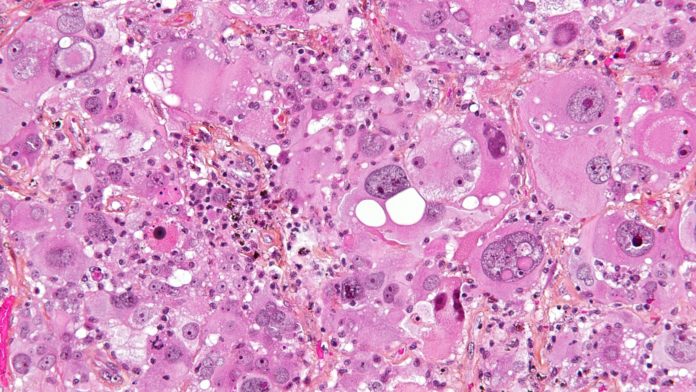
Glioblastoma Multiforme (GBM) is a highly aggressive form of brain cancer with limited options for treatment and poor survival rates. It’s a leading cause of cancer-related deaths among young children and teens, and also took the life of Tragically Hip frontman Gord Downie.
In a new study, Canadian researchers have effectively “reverse engineered” GBM stem cells sourced from patients and discovered several potential targets for treatment.
The collaborative study by researchers from the Universities of Calgary and Toronto is the first to “systematically profile a large panel of patient-derived brain tumour cells that have stem cell properties,” according to an accompanying press release.
“We think that, in one big experiment, we have uncovered many new targets for glioblastoma, some of which were surprising,” says Peter Dirks, a co-principal investigator.
“These GBM stem cells are also resistant to treatment, which is one reason that these tumours are so hard to cure. We need new ways to disrupt these cells specifically if we are going to give people a better chance of survival.”
Reverse engineering GBM stem cells to locate the cancer’s ‘engine’
Cancer stem cells drive the development of the disease, so to target these cells using drugs, scientists need to know which genes are responsible for directing growth.
Researchers created GBM stem cell cultures sourced from 10 patients and analyzed all 20,000 genes from every sample to assess which are important for the proliferation of cancer cells. Their analysis was completed using CRISPR-Cas9, a powerful gene editing tool that allows scientists to edit parts of the genome by reconfiguring sections of the DNA sequence.
Co-principal investigator Stephane Angers compared the Cas9 protein element of this technology to a homing device which tracks down what genes are important.
“It’s like you take all the components to make a car and put them on the sidewalk,” says Angers. “An engineer could tell you which pieces of the engine are important to make the car move forward. Our study basically was able to identify the pieces of the engine, which are the pieces within the cells that are really important to make the tumour cell grow.”
Under the direction of Angers, the team found around 400 genes that may be significant for the development of the disease.
Especially notable was a gene called DOT1L which was found to be necessary for tumour persistence in 7 out of 10 cultures. A collaboration headed by Samuel Weiss, a professor from the University of Calgary, was able to show in preclinical models that a drug used to tackle leukaemia could block the DOT1L gene used in GBM stem cells.
“This is promising because it uncovered a biological process, not previously suspected to be implicated in [GBM], for which a small molecule drug already exists,” says Angers.
Additional research may explain failure of existing treatments
A separate experiment by the team explored which genes are involved in the resistance to temozolomide chemotherapy (TMZ therapy) using CRISPR-Cas9 screens. TMZ therapy was the last major step forward in terms of treatment for GBM, but its efficacy is limited and can even make matters worse by providing tumour cells with more mutations.
This experiment highlighted genes involved in a biochemical network known as the Fanconi anemia pathway which has a hand in several processes such as DNA repair and replication. Previous research has also shown this network to be a point of resistance to TMZ.
Moving forward, Angers commented that the next step is to start targeting the suspected genes with tailored drugs and other tools. If successful, GBM patients may finally have a worthy defence against this devastating disease.
“We and others are now going to try and develop inhibitors, drugs, small molecules, antibodies to interrupt the function of these genes,” says Angers.
“What we’re now providing the scientific community is the blue book, if you will, the instruction manual, of how our GBM cancer cells are wired and what is important in their circuits for driving the growth programs.”