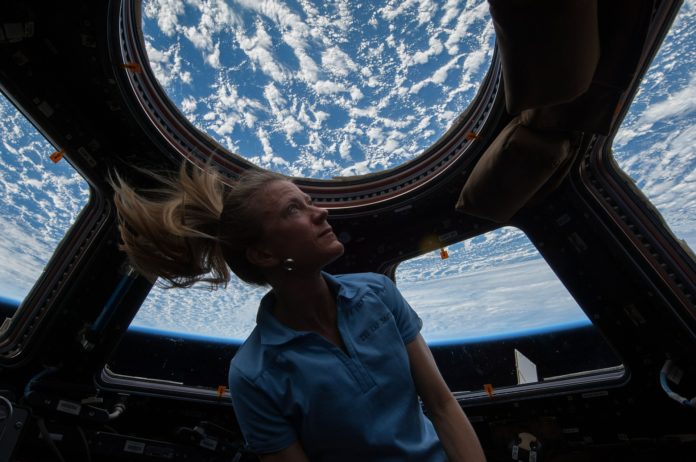
Since the launch of the International Space Station (ISS) in 1998, astronauts from around the world have been responsible not just for performing scientific experiments in outer space, but also for making sure the interior of the ISS remains clean — and safe. Any contaminated surfaces could spell danger for the astronauts on board.
Now, a team of scientists from McGill University and the University of Montreal have found a way to make identifying microbial health hazards a lot easier.
Their study focuses on a new method to identify and map different microbial species aboard the ISS, and was published in Environmental Microbiology.
The trouble with traditional identification techniques
Although scientists have long known that the ISS is teeming with bacteria — the station is airtight, after all, with almost no chance for bacteria brought in to escape — relatively little was known about the different types of microbes that call the ISS home.
Previous studies into the microbiome aboard the ISS had used a traditional bacterial identification method that made use of the so-called 16S ribosomal RNA (rRNA) gene. This gene is present in all bacteria, and acts like a barcode that scientists can use to distinguish between different species within the same genus. In complex environments, however, wide variation can exist between different copies of the same gene in an organism. In these cases, the method isn’t as reliable.
To more robustly and accurately identify microbial species aboard the ISS, the team made use of a new method called ANCHOR. Unlike the traditional 16S rRNA gene, this new technique uses paired-end sequencing: a technique that allows for both ends of an RNA or DNA fragment to be analyzed. ANCHOR also combines multiple high-complexity samples, as well as multiple reference databases, to make the most accurate bacterial identification possible.
To test their new method, the researchers applied it to several artificial datasets, as well as surface swabs taken from various parts of the ISS. The ISS sample was specifically chosen because it was, as the authors described in their paper, both “technically nonideal and biologically complex.”
Microbiome aboard the ISS is more diverse than expected
The results from the ANCHOR method not only agreed with previous studies of the ISS dataset, but also revealed much more about this complex outer space environment than was previously known. In particular, ANCHOR was able to identify 714 unique bacterial species thriving throughout the space station.
The technique was also able to map differences between the sleeping quarters, which were dominated by human gastrointestinal bacteria, and the laboratory, which was dominated by bacteria from non-human research animals that had been aboard the station in the past.
“Scientists have a well-documented understanding of broad bacterial families on the ISS,” explained Emmanuel Gonzalez, a bioinformatics consultant at McGill University and the lead author of the study, “however, we’ve discovered a more diverse bacterial ecosystem [than] we ever expected.”
The authors are currently investigating how this method can be used not just in outer space, but in a range of complex environments right here on Earth.
“The new methodology provides us spectacular snap-shots of the bacterial world in space,” said co-author Nicholas Brereton, a researcher at the University of Montreal’s Institut de recherche en biologie végétale.
“The possibilities of applying this method to explore new microbiome environments are really exciting.”